Topics in language and cognition

- What is Language?
- Professor Marc Hauser defines language as an internal process that involves ways of manipulating symbols. It is qualitatively different to "communication."
- SOURCE: G2C
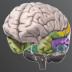
- Temporal Lobe
- The temporal lobes contain a large number of substructures, whose functions include perception, face recognition, object recognition, memory, language, and emotion.
- SOURCE: G2C

- Chromosome 7: FOXP2 gene involved in language, Matt Ridley
- Matt Ridley talks about chromosome 7, FOXP2 gene involved in language.
- SOURCE: DNAi

- Chimps, humans, and language, Svante Paabo
- Evolutionary geneticist Svante Paabo speaks about the differences between chimp and humans and a genetic basis for language.
- SOURCE: DNAi

- Broca's Area - Primary Functions
- Dr. Sukhi Shergill discusses the importance of Broca's area to generating speech.
- SOURCE: G2C

- Language and Animals
- Professor Marc Hauser discusses the differences and similarities between humans and primates in terms of language ability.
- SOURCE: G2C

- Do Autism Symptoms Cluster Together?
- Professor David Skuse explains that the language and social difficulties associated with autism correlate more closely than repetitive behavior symptoms.
- SOURCE: G2C

- Social cognition (2)
- Doctor Thomas Insel defines social cognition as the way we process information about recognition, social memory, social motivation, and language.
- SOURCE: G2C
Videos about linkage analysis:

- Linkage studies versus quantitative genetics
- Professor Allen Moore outlines the differences between quantitative genetics and linkage studies. With quantitative genetics it is not necessary to begin with the physical DNA.
- SOURCE: G2C
- transcript: So linkage studies and quantitative genetics both rely on statistical associations, but linkage is actually taking a physical piece of the genome and showing that that physical piece is associated with differences in a given phenotype. So if you have differences in the sequence, so you have sequenced everything again, and you find that those differences correspond to differences in a trait that you are interested in then you can associate those two and show that there are regions of particular genome that are influencing differences in the phenotype of interest. Whereas quantitative genetics, you don’t have to know anything about the genetics; you actually start with the trait itself and then you use, whether or not individuals that are related are more similar in those traits then unrelated. So quantitative genetics is somewhat easier because it is a little more indirect, we don’t have to have the information on the genome in order to use quantitative genetics, so that’s a first step. The second step then is to associate differences in genomes with the differences in the traits.

- Linkage versus association studies
- Professor Allen Moore describes the differences between linkage and association studies, which are low- and high-resolution techniques used to search for candidate genes.
- SOURCE: G2C
- transcript: Linkage is actually looking at physical segments of the genome that are associated with given traits. Association studies go from the other direction, saying, ‘given different pieces of the genome, can we then look for different traits that are associated with those different segments of genome?’ So we know that individuals don’t have the same genetic makeup. They have the same DNA, but the DNA has different sequences or is expressed differently, and that’s what causes differences among different individuals. So the question is that if we have a trait, particularly a disease trait, can we find and associate that with differences among individuals in the population? So a linkage study is just saying, ‘can we say that there is an association between pieces of the DNA and a trait of interest?’ Association studies are saying, ‘what are the differences we see?' in order to find differences in the traits, particularly disease traits, among different individuals.

- Linkage versus Association Studies (video is not provided yet)
- Doctor Anil Malhotra compares (older) linkage and (more modern) association techniques for identifying candidate genes for disorders.
- SOURCE: G2C
- transcript: Linkage analysis was a very popular method for detecting genes of major effect particularly in psychiatry. It was used mostly in the '80s and perhaps early 1990s usually based on within-family design either sibling pairs or large multiplex pedigrees. As I said, it is really optimally designed for disorders in which their genes have major effect. One of the things I think that came out of that generation of linkage studies was [that] it is relatively clear that if there are genes of major effect they are relatively rare and relatively isolated populations within schizophrenia. Association studies take the opposite approach. In general, there are case-control association studies, though there are family-based approaches also, which hopefully are going to find genes that have less of a strong effect and thus maybe multiple genes of lesser effect can be detected. With the association approach, where we essentially compare allele frequencies between cases and controls and then examine whatever number of genes or number of polymorphisms in individual study.

- Linkage and association studies
- Professor James Potash describes the difference between linkage and association studies, which are two ways of locating candidate genes. These are discussed in reference to bipolar disorder.
- SOURCE: G2C
- transcript: There are two major approaches to looking for candidate genes in bipolar disorder; the one that has been focused on the most is the gene-mapping approach, which is a hypothesis-free approach. Part of the reason that’s so attractive is because there has been so little that we’ve understood about the pathophysiology of the illness, of the way the illness unfolds in the brain, that we’ve thought that if we can from a hypothesis-free approach go directly to the gene it would be a huge advance. Linkage was the original way to go about that; that was a very low resolution way to screen the genome, looking for regions where there might be bipolar disorder genes. We are now at a much higher resolution approach that’s referred to as whole genome association, where you can look at seven hundred thousand to a million different places in the genome all at once; that’s the state of the art right now.
-
